
Department of Cell and Developmental Biology
University of California San Diego
RESEARCH
Our lab studies the molecular and cell biological mechanisms of immune checkpoint receptors (ICRs) — inhibitory receptors that act as molecular brakes on the immune system. These pathways are essential for preventing autoimmunity, but they can also be exploited by tumors and pathogens to suppress immune responses. Although drugs targeting immune checkpoints such as PD1, CTLA4, and LAG3 have transformed cancer therapy, they benefit only a subset of patients and often cause immune-related toxicities. A deeper mechanistic understanding of these receptors is needed to expand the reach and precision of immunotherapy.
Our research integrates biophysical, biochemical, cell biological, and immunological approaches to investigate immune checkpoint function across multiple scales, from reconstituted molecular systems to cells and animal models. Our work is currently organized around three main themes.
RESEARCH AREAS
1. Signal transduction mechanisms of immune checkpoint receptors
A central goal of the lab is to understand how immune checkpoint receptors transmit inhibitory signals inside cells. We are particularly interested in how receptors such as PD1, CTLA4, and related pathways engage downstream effectors and reorganize signaling networks that control T cell activation, differentiation, and anti-tumor function. Our long-term aim is to define the core molecular logic of inhibitory signaling and use that knowledge to guide more effective therapeutic strategies.
We reported that PD1 and BTLA inhibitory signaling are only partially dependent on Shp2 and its paralog Shp1, as both receptors retain substantial inhibitory activity in Shp1/Shp2 double-knockout T cells (PMID: 32437509). Together with findings from other groups, this work suggests that PD1 signaling also involves Shp1/2-independent mechanisms.
In recent work (https://doi.org/10.64898/2026.03.09.710629), we found that ligand-bound PD1 can trigger Shp2 condensation, organizing inhibitory signalosomes through selective substrate partitioning. This work suggests that Shp2 liquid–liquid phase separation (LLPS) is an intrinsic organizing principle of the PD1 pathway. Mechanistically, PD1-induced Shp2 condensation promotes the co-compartmentalization of signaling substrates such as CD3ζ and CD28, facilitating their dephosphorylation and suppressing T cell activation. We also found that mutations that selectively disrupt Shp2 self-association weaken PD1 microcluster formation and impair inhibitory function. Together, these findings support a model in which enzymatic activation, substrate selectivity, and mesoscale assembly are tightly integrated in immune checkpoint signaling.
Going forward, we are combining proteomics, membrane reconstitution, cellular imaging, and functional immunology to dissect the signaling mechanisms of PD1 and other immune checkpoint receptors in greater detail.
2. Regulation of immune checkpoints through cell-intrinsic cis-interactions
A major focus of our lab is how immune receptor function is regulated by cis-interactions between receptors and ligands on the same cell membrane. Our work has uncovered several such pathways, including PD-L1:CD80 (PMID: 31757674), B7:CD28 (PMID: 37160118), and B7:CTLA4 (PMID: 37042938), and defined their roles in tuning T cell signaling. We further found that CTLA4 limits trogocytosis-mediated self-stimulation by depleting acquired ligands in cis, revealing a cell-intrinsic mechanism that extends beyond the classical APC–T cell interface. Together, these studies identify cis-interactions as a new conceptual layer in immunoreceptor signaling.
Going forward, we aim to define how broadly these cell-intrinsic interactions operate across immune receptor systems and how they can be leveraged to modulate immunity more precisely in disease and therapy.
3. Cross-species divergence of immune checkpoint receptors
Most of what we know about immune checkpoint biology comes from mouse models, yet many checkpoint receptors are only moderately conserved between mouse and human. Our lab is increasingly interested in how such evolutionary divergence affects mechanism, function, and therapeutic response.
In recent work (PMID: 39752535), we carried out a systematic comparison of human and mouse PD1 and found that human PD1 is more inhibitory than its mouse counterpart. This difference arises from stronger interactions with both PDL1/2 and the effector phosphatase Shp2. We further identified a conserved motif present across vertebrate PD1 orthologs but absent in rodents that helps explain this difference. Functional studies showed that replacing the intracellular domain of mouse PD1 with that of human PD1 profoundly alters T cell behavior and response to anti-PD1 therapy. These findings highlight important species-specific features of the PD1 pathway and raise broader questions about how faithfully mouse models capture the biology of human immune checkpoints.
Going forward, we aim to extend this evolutionary and mechanistic framework to other checkpoint receptors, many of which also show substantial divergence between mouse and human orthologs.
APPROACHES
1. Reconstitution of intracellular signaling: We directly and quantitatively study T cell signaling reactions using membrane reconstitution assays, in which purified proteins are reconstituted to synthetic lipid bilayers. In membrane reaction kinetics and net output of signaling modules, can be probed by fluorescence and microscopy readouts.

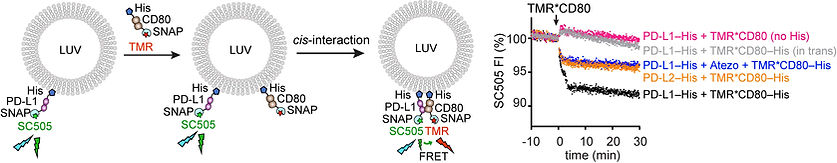
2. Reconstitution of cis versus trans-interactions: We developed membrane reconstitution assays to detect and characterize either cis or trans-interactions among T cell surface proteins.
3. Live cell imaging of receptor dyanmics: We visualize the spatiotemporal dynamics of immune checkpoints using live T-cell : bilayer hybrids, in which the APCs are replaced with a ligands-containing supported lipid bilayer.

4. Proteomic analysis of immune checkpoint interactomes: We discover new interactors of immune checkpoints using quantitative proteomics.
5. T : APC co-cultures & animal models: We cross-check our findings in conventional T cell signaling assays in both ex vivo cultures and in vivo models.


